This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License
using CovidSEIR, Plots, DataFrames, JLD2, StatsPlots, Dates, MCMCChains
Plots.pyplot()
jmddir = normpath(joinpath(dirname(Base.find_package("CovidSEIR")),"..","docs","jmd"))
covdf = covidjhudata();
United States¶
us = CountryData(covdf, "US")
CovidSEIR.CountryData{Float64,Int64}(3.2716743e8, [1, 2, 3, 4, 5, 6, 7, 8,
9, 10 … 77, 78, 79, 80, 81, 82, 83, 84, 85, 86], [0.0, 0.0, 0.0, 0.0, 0.0
, 0.0, 0.0, 0.0, 0.0, 0.0 … 12722.0, 14695.0, 16478.0, 18586.0, 20462.0,
22019.0, 23528.0, 25831.0, 28325.0, 32916.0], [0.0, 0.0, 0.0, 0.0, 0.0, 0.0
, 0.0, 0.0, 0.0, 0.0 … 21763.0, 23559.0, 25410.0, 28790.0, 31270.0, 32988
.0, 43482.0, 47763.0, 52096.0, 54703.0], [1.0, 1.0, 2.0, 2.0, 5.0, 5.0, 5.0
, 5.0, 5.0, 7.0 … 361738.0, 390798.0, 419549.0, 449159.0, 474664.0, 50030
6.0, 513609.0, 534076.0, 555929.0, 580182.0])
using Turing
mdl = CovidSEIR.TimeVarying.countrymodel(us)
cc = Turing.psample(mdl, NUTS(0.65), 5000, 4)
import JLD2
JLD2.@save "$jmddir/us_tv_$(Dates.today()).jld2" cc
JLD2.@load "$jmddir/us_dhmc_2020-04-13.jld2" cc;
cc = MCMCChains.Chains(collect(cc.value.data), replace.(cc.name_map.parameters, r"([^\[])([1-9])" => s"\1[\2]"))
Object of type Chains, with data of type 5000×15×4 Array{Float64,3}
Iterations = 1:5000
Thinning interval = 1
Chains = 1, 2, 3, 4
Samples per chain = 5000
parameters = τ, sigD, sigC, sigRc, a, pE0, p[1], p[2], β[1], β[2], β
[3], γ[1], γ[2], ρ[1], ρ[2]
2-element Array{MCMCChains.ChainDataFrame,1}
Summary Statistics
parameters mean std naive_se mcse ess r_hat
────────── ───────── ─────── ──────── ────── ──────── ──────
τ 0.0001 0.0000 0.0000 0.0000 81.9924 1.2819
sigD 137.7127 8.0087 0.0566 0.3734 140.4813 1.0810
sigC 1334.2734 92.6313 0.6550 4.5797 97.6593 1.2625
sigRc 731.7612 43.2561 0.3059 2.2038 96.7621 1.2997
a 0.7609 0.1862 0.0013 0.0131 80.3213 3.9037
pE0 0.0000 0.0000 0.0000 0.0000 100.5049 1.1776
p[1] 0.0193 0.0021 0.0000 0.0001 80.3213 1.7318
p[2] 0.0045 0.0000 0.0000 0.0000 141.0271 1.0764
β[1] 0.8113 1.2418 0.0088 0.0876 80.3213 9.1362
β[2] 1.1373 0.6511 0.0046 0.0422 80.3213 1.5722
β[3] 3.9270 0.7566 0.0053 0.0529 80.3213 3.8765
γ[1] 2.9379 0.1078 0.0008 0.0076 80.3213 7.4234
γ[2] 0.0067 0.0003 0.0000 0.0000 80.3213 1.4663
ρ[1] 0.2486 0.0306 0.0002 0.0021 80.3213 2.2939
ρ[2] 61.2539 0.4053 0.0029 0.0241 80.3213 1.3951
Quantiles
parameters 2.5% 25.0% 50.0% 75.0% 97.5%
────────── ───────── ───────── ───────── ───────── ─────────
τ 0.0000 0.0000 0.0000 0.0001 0.0002
sigD 118.6077 135.2341 139.7070 141.1686 152.2954
sigC 1155.0157 1294.3130 1305.6873 1394.1663 1504.5102
sigRc 653.7688 701.9086 724.4422 762.5418 816.5018
a 0.3665 0.5791 0.7621 0.9520 0.9683
pE0 0.0000 0.0000 0.0000 0.0000 0.0000
p[1] 0.0151 0.0178 0.0190 0.0216 0.0223
p[2] 0.0045 0.0045 0.0045 0.0045 0.0045
β[1] 0.0000 0.0000 0.1534 0.7802 3.2127
β[2] 0.0723 0.7875 1.1188 1.5326 2.6644
β[3] 2.4007 4.1450 4.1807 4.4108 4.7409
γ[1] 2.7158 2.8516 3.0000 3.0000 3.0000
γ[2] 0.0061 0.0064 0.0067 0.0070 0.0070
ρ[1] 0.2132 0.2236 0.2414 0.2667 0.3312
ρ[2] 60.5702 60.9084 61.1496 61.5801 62.0245
Estimates¶
plot(cc)
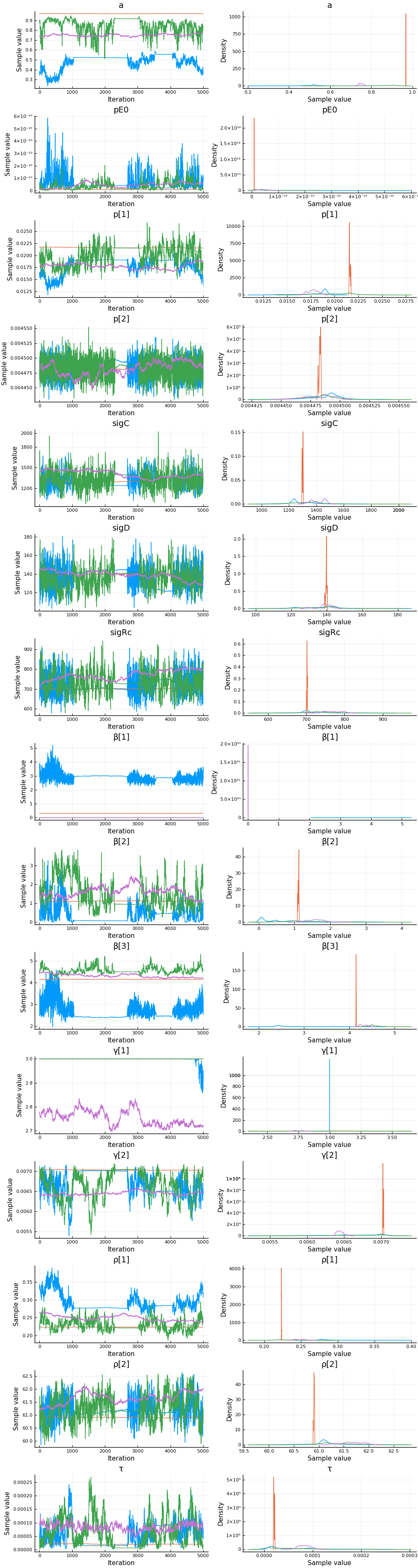
describe(cc)
2-element Array{MCMCChains.ChainDataFrame,1}
Summary Statistics
parameters mean std naive_se mcse ess r_hat
────────── ───────── ─────── ──────── ────── ──────── ──────
τ 0.0001 0.0000 0.0000 0.0000 81.9924 1.2819
sigD 137.7127 8.0087 0.0566 0.3734 140.4813 1.0810
sigC 1334.2734 92.6313 0.6550 4.5797 97.6593 1.2625
sigRc 731.7612 43.2561 0.3059 2.2038 96.7621 1.2997
a 0.7609 0.1862 0.0013 0.0131 80.3213 3.9037
pE0 0.0000 0.0000 0.0000 0.0000 100.5049 1.1776
p[1] 0.0193 0.0021 0.0000 0.0001 80.3213 1.7318
p[2] 0.0045 0.0000 0.0000 0.0000 141.0271 1.0764
β[1] 0.8113 1.2418 0.0088 0.0876 80.3213 9.1362
β[2] 1.1373 0.6511 0.0046 0.0422 80.3213 1.5722
β[3] 3.9270 0.7566 0.0053 0.0529 80.3213 3.8765
γ[1] 2.9379 0.1078 0.0008 0.0076 80.3213 7.4234
γ[2] 0.0067 0.0003 0.0000 0.0000 80.3213 1.4663
ρ[1] 0.2486 0.0306 0.0002 0.0021 80.3213 2.2939
ρ[2] 61.2539 0.4053 0.0029 0.0241 80.3213 1.3951
Quantiles
parameters 2.5% 25.0% 50.0% 75.0% 97.5%
────────── ───────── ───────── ───────── ───────── ─────────
τ 0.0000 0.0000 0.0000 0.0001 0.0002
sigD 118.6077 135.2341 139.7070 141.1686 152.2954
sigC 1155.0157 1294.3130 1305.6873 1394.1663 1504.5102
sigRc 653.7688 701.9086 724.4422 762.5418 816.5018
a 0.3665 0.5791 0.7621 0.9520 0.9683
pE0 0.0000 0.0000 0.0000 0.0000 0.0000
p[1] 0.0151 0.0178 0.0190 0.0216 0.0223
p[2] 0.0045 0.0045 0.0045 0.0045 0.0045
β[1] 0.0000 0.0000 0.1534 0.7802 3.2127
β[2] 0.0723 0.7875 1.1188 1.5326 2.6644
β[3] 2.4007 4.1450 4.1807 4.4108 4.7409
γ[1] 2.7158 2.8516 3.0000 3.0000 3.0000
γ[2] 0.0061 0.0064 0.0067 0.0070 0.0070
ρ[1] 0.2132 0.2236 0.2414 0.2667 0.3312
ρ[2] 60.5702 60.9084 61.1496 61.5801 62.0245
Fit¶
sdf = simtrajectories(cc, us, 1:200)
f = plotvars(sdf, us)
plot(f.fit, ylim=(0, maximum(us.active)*1.3))
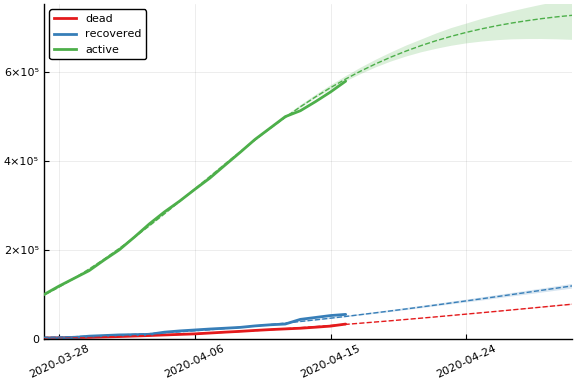
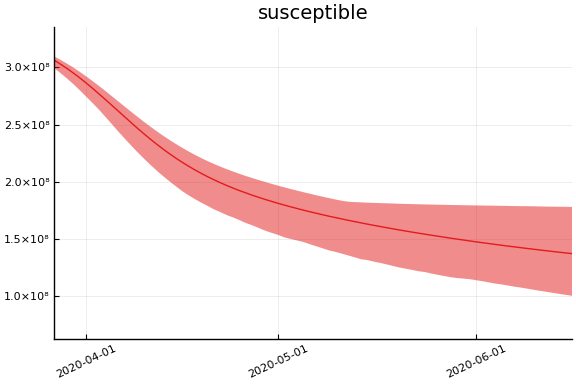
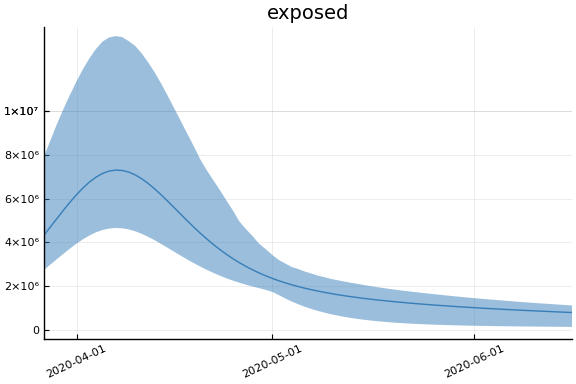
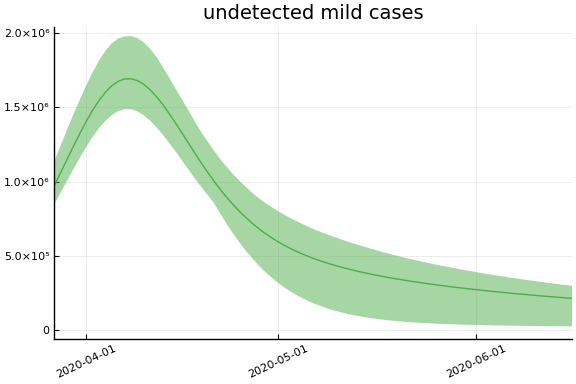
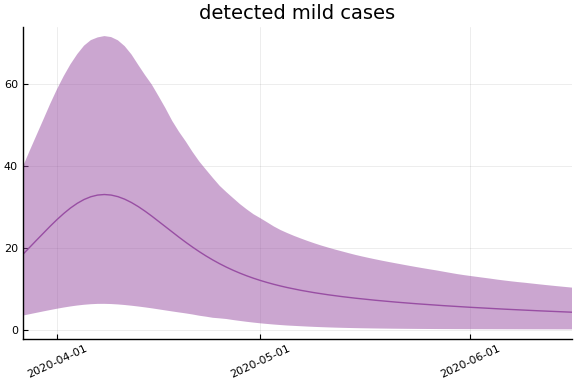
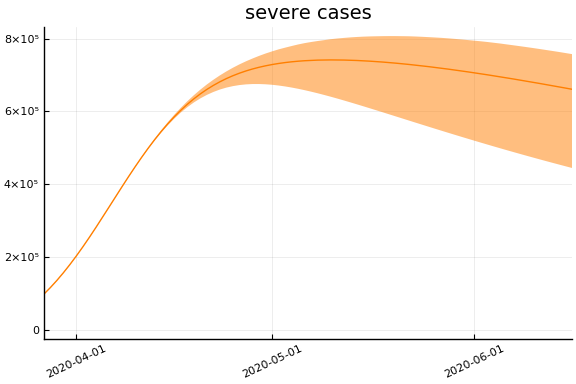
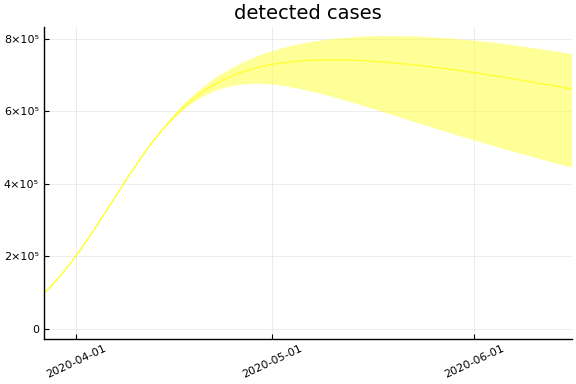
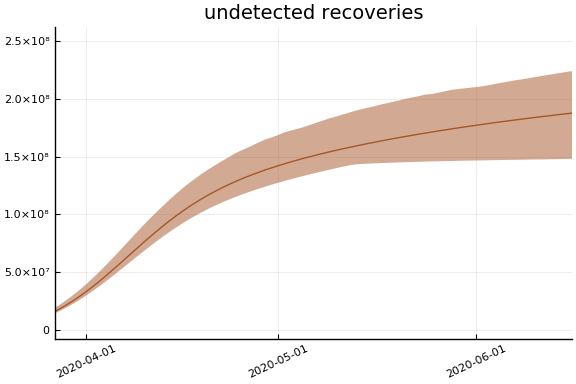
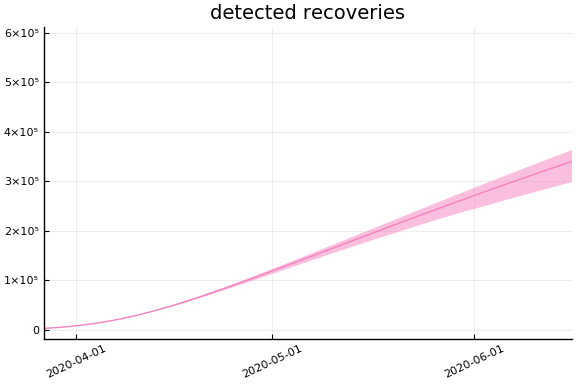
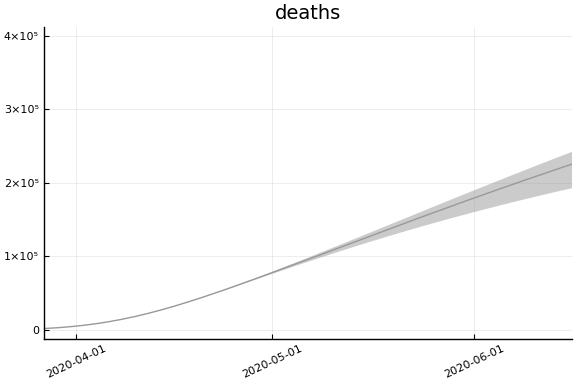
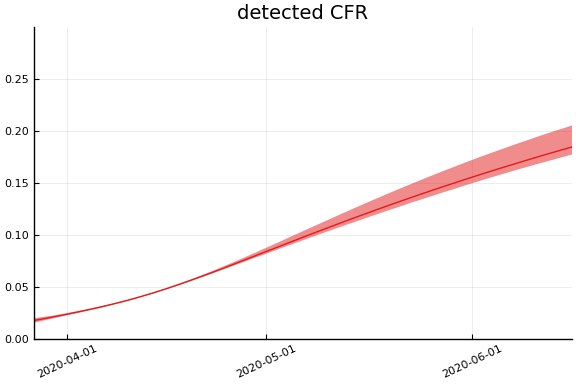
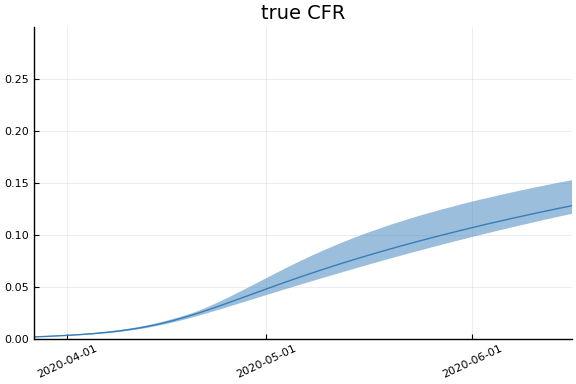
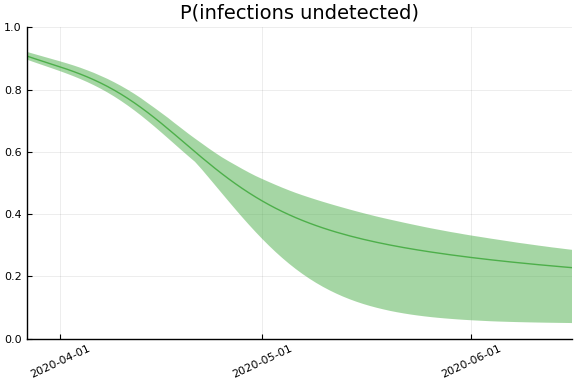
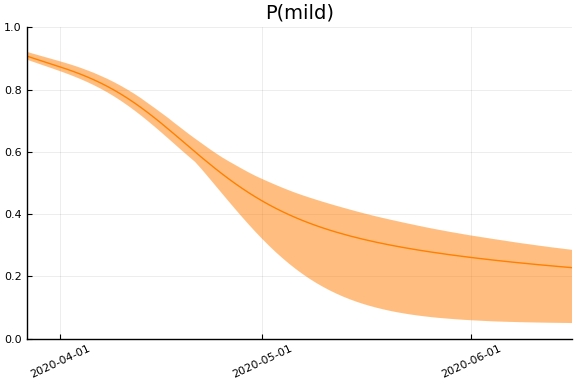
Implications¶
for fig in f.trajectories
display(plot(fig))
end